Serial cloner for windows6/22/2023 
Users can also create their own custom databases from pdb files in addition to the default database, and this is most useful when the proteins of interest are similar to the proteins used to make the database, for example from pdb files of the same protein family, protein location (membrane/cytoplasm) or environment (halophilic, mesophilic, thermophilic). The default database of patterns was generated from publicly available pdb structures. The custom protein secondary structures prediction algorithm is based on patterns analysis of alpha helices, beta strands and turns. These custom algorithms are faster than the Needleman-Wunsch algorithm for large sequences having high homology. In contrast to the Needleman-Wunsch algorithm, the custom algorithms provide correct alignment independent of the size of the sequences indels. The program includes custom algorithms for: the protein/DNA sequences alignment, protein secondary structures prediction, RNA-seq differential expression. Analysis of Gel/Blot photo and calculation of relative fold-change in selected Gel/Blot bands/spots.Calculators for: Dilution, Molarity, Ligation, SDS PAG, Ammonium sulfate, Bacteria growth.RNA-Seq DE: genes and miRNAs differential expression.Find/highlight protease sites and protein fragments.Prediction of protein secondary structures.Multiple alignment and realignment of DNA/RNA/protein sequences.Find/highlight DNA restriction sites and DNA fragments.Copy/paste the selected DNA sequence with features.Generate DNA features database and perform features annotation.Generate and edit DNA/protein features for genbank/genpept format files.Generate Plasmid, Linear and Text map using feature annotations from genbank file.View GC content, CpG islands and Stem Loops.View ABI sequencing chromatogram, edit sequence.Find sequence motifs (repeats, palindromes, stem loops and arbitrary sequences) in DNA sequence.Tm, %GC, Mw, length of selected DNA and protein Mw (average and monoisotopic), length of protein, amino acid composition, protein net charge.Transcribe, reverse complement and translate DNA sequence.The program reads files created by Serial Cloner(.xdna), DNA Strider(.xdna), SnapGene(.dna) and ApE (.ape) programs. The user interface was developed around a minimalistic design and ease of use, such as: direct access to functions in less than three clicks, essential functionality with a minimum of settings, minimisation of windows (uncluttered screen). It also provides a set of useful calculators for molecular biology. It is a suite of the bioinformatic tools for the analysis of DNA, RNA, and protein sequences, as well as images of gels, blots, cells and colonies. This program is aimed at researches in the molecular biology field for use in their everyday routine work. BioLabDonkey is one time purchase app with lifetime updates including new features. While we have not verified the apps ourselves yet, our users have suggested 3 different NUCL openers which you will find listed below.BioLabDonkey is a program for macOS. Important: Different programs may use files with the NUCL file extension for different purposes, so unless you are sure which format your NUCL file is, you may need to try a few different programs. 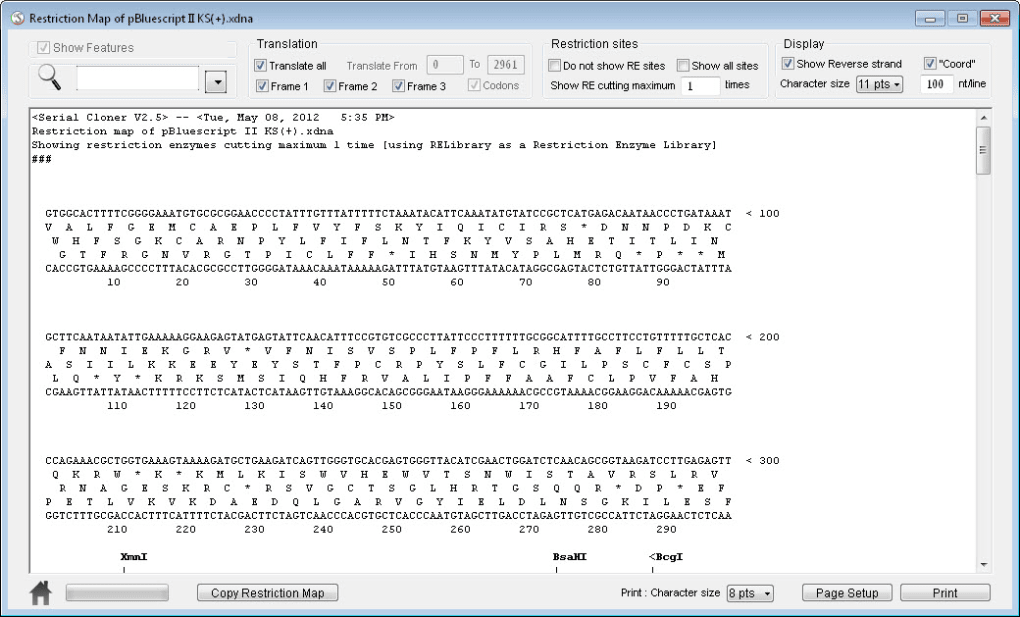
We are continually working on adding more file type descriptions to the site, so if you have information about NUCL files that you think will help others, please use the Update Info link below to submit it to us - we'd love to hear from you! How to open NUCL files  
While we do not yet describe the NUCL file format and its common uses, we do know which programs are known to open these files, as we receive dozens of suggestions from users like yourself every day about specific file types and which programs they use to open them. However, different programs may use the NUCL file type for different types of data. The NUCL file extension indicates to your device which app can open the file.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
